 Īny additional information required to reanalyze the data reported in this paper is available from the lead contact upon request.Īn original code used for determining ploidy (gene copy number) and allele genotype of KIR is available at with reference data curated from IPD-KIR allele database, both of which are publicly available ( ).read.GenoDive imports data from text files formatted for this. Make the file input display none and boom, it works in IE9+ seamlessly. A much better way to do this is to just create a file input and a label that links to it. Other web resources used in this study are listed in the key resources table. GenoDive is a Mac-only program for population genetic analysis that allows for polyploid data. Works on every other type of element as expected, but doesn't work on file inputs. Accession numbers are listed in the key resources table. The genotype data of BBJ used in this study are available from the Japanese Genotype-phenotype Archive (JGA) through application at. The International Histocompatibility Working Group (IHWG) cell lines were obtained from ECACC HLA typed collection ( ). One major feature of genodive is that it supports both diploid and polyploid data, up to octaploidy (2n 8x) for some analyses, but up to hexadecaploidy (2n 16x) for other analyses. The genomic DNA we used for KIR sequencing were obtained from a cell line established by the Japan Biological Informatics Consortium (JBIC), and can be purchased at. Furthermore, genodive seamlessly supports 15 different file formats for importing or exporting data from or to other programs. read.GenoDive imports data from text files formatted for this program. Accession numbers are listed in the key resources table. GenoDive is a Mac-only program for population genetic analysis that allows for polyploid data. Individual-level KIR alleles and KIR imputation reference panel used in this study are deposited at the National Bioscience Database Center (NBDC) Human Database ( ) and are publicly available as of the date of publication.  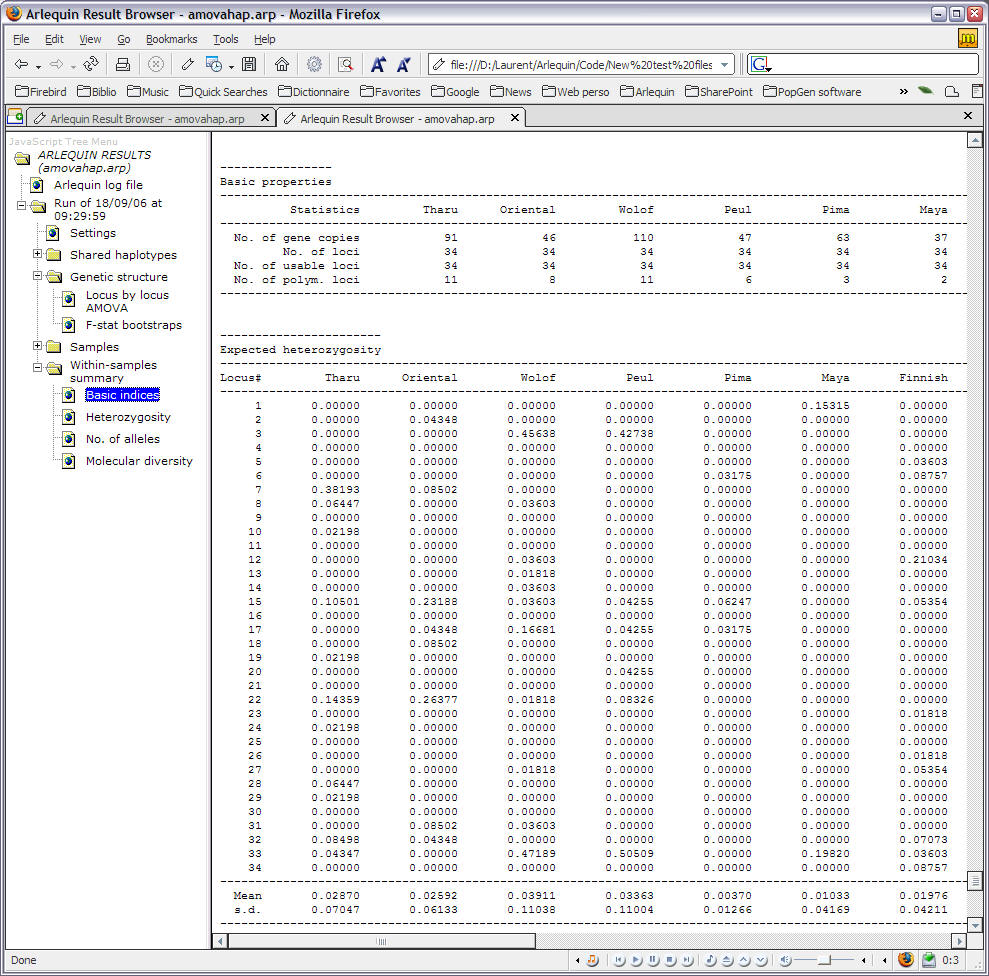
Our pipeline presents a broadly applicable framework to evaluate innate immunity in large-scale datasets. We observe a dearth of genome-wide significant associations, even in immune traits implicated previously to be associated with KIR (the smallest p = 1.5 × 10 −4). We construct a KIR imputation reference panel (n reference = 689, imputation accuracy = 99.7%), apply it to biobank genotype (n total = 169,907), and perform phenome-wide association studies of 85 traits. We define 118 alleles in 13 genes and demonstrate a linkage disequilibrium structure within and across KIR centromeric and telomeric regions. We devise a bioinformatics pipeline incorporating copy number estimation and insertion or deletion (indel) calling for high-resolution KIR genotyping. Here we conduct deep sequencing of 16 KIR genes in 1,173 individuals. Despite its role in immunity, the complex genomic structure has limited a deep understanding of the KIR genomic landscape. From what I learned, sample sizes of n=1 can never be significant.Īlso I calculated Fst values with GenoDive, wich is leading to slightly different Fst values (mean diference 0.0151, sd=0.0699).The killer cell immunoglobulin-like receptor (KIR) recognizes human leukocyte antigen (HLA) class I molecules and modulates the function of natural killer cells. We also encourage you to check the files with your own antivirus before launching the installation. Thank you for downloading GenoDive for Mac from our software library The software is periodically scanned by our antivirus system. 
How is that possible? I did check the input file, no problem there. Download GenoDive Free If your download is not starting, click here. Fst values of "A" compared to other populations shows significant p-values (p<0.05). To define a file-select field that allows multiple files to be selected, add the multiple attribute. However I have a population "A" with only one individual. The defines a file-select field and a 'Browse' button for file uploads.Usage read.GenoDive (infile) Arguments Details GenoDive is a Mac-only program for population genetic analysis that allows for polyploid data. I did calculate Fst values with 3 microsatellite makers for 29 populations using Arlequin 3.5.1.3 (Population pairwise Fst values: Compute F-statistics on haplotype frequencies only). read.GenoDive takes a text file in the format for the software GenoDive and produces a genambig object. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
